HOME > Program Members > Hideyuki Saya

Hideyuki Saya

Professor, Division of Gene Regulation, Institute for Advanced Medical Research, Graduate School of Medicine, Keio University
Hideyuki Saya, MD, PhD
hsaya@a5.keio.jp
http://genereg.jp/index.html
Theme
Tumors are thought to contain a small population of cells designated as cancer stem cells (CSCs), that possess both the ability for self-renewal and the ability to differentiate into multiple lineages, that are considered to play key roles in the initiation and maintenance of cancer. Moreover, CSCs are known to be resistant to radiotherapy and chemotherapy. Therefore, intensive characterization of CSCs is extremely important for the development of new effective therapeutic approaches. In addition to the cancer stem cell theory it is the very important concept that epithelial mesenchymal transition (EMT) plays a key role in cancer invasion and metastasis. The goal of our project is to understand the mechanisms underlying the genesis and maintenance of CSCs and EMT in cancer cells. The specific approaches we propose to use to reach this goal are: i) to identify the CSC fraction in mouse cancer models developed by genetic manipulation and to analyze the molecular mechanism underlying the switch between CSCs and non-CSCs, and ii) to develop a new cell-based assay system to evaluate EMT in vitro and to identify the molecules regulating the induction and suppression of EMT. Both approaches will also be applied for drug screening.
Research activities
Characterization and regulation of CSCs
We have established mouse malignant tumor models by minimum essential genetic manipulations (Fig. 1). We have developed a mouse model of pre-B lymphoblastic leukemia/lymphoma or osteosarcoma. We have identified the cellular fraction in these tumors that is tumorigenic as well as resistant to chemotherapeutic agents. We are currently analyzing the differences between the resistant cells and sensitive cells in order to elucidate the molecular mechanism regulating the switch.
To visualize the cell cycle progression and cell movements in a live vertebrate, we have developed the transgenic histone H2B-GFP Medaka line. The system has high specificity, with strong GFP labeling of the chromatin architecture (Cell Cycle, 2009).
Identification of signals regulating EMT and the development of EMT-inhibitors:
We have established a system that allows assessment of EMT induction in vivo. This assay system enabled us to identify the key signal cascades for the induction and maintenance of EMT and was applied to screen chemical compounds to control EMT.
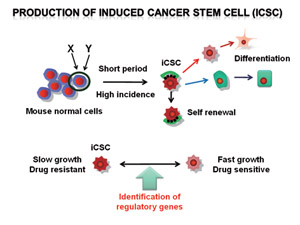
Fig.1 Production and characterization of induced cancer stem cells (iCSC) and development of new strategies against CSCs.
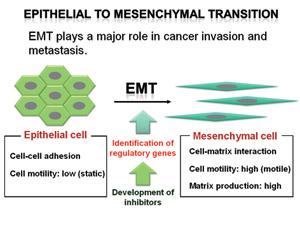
Fig.2 Identification of the molecular events regulating EMT in cancer cells and development of a new therapeutic approach to suppress EMT.
Selected Paper
- Sampetrean O, Iida S, Makino S, Matsuzaki Y, Ohno K, Saya H: Reversible whole-organism cell cycle arrest in a living vertebrate. Cell Cycle 8(4): [Epub ahead of print], 2009
- Arima Y, Inoue Y, Shibata T, Hayashi H, Nagano O, Saya H and Taya Y: Rb depletion results in deregulation of E-cadherin and induction of cellular phenotypic changes that are characteristics of the epithelial to mesenchymal transition. Cancer Res 68: 5104-5112, 2008.
- Takahashi A, Ohtani N, Yamakoshi K, Iida S, Tahara H, Nakayama K, Nakayama KI, Ide T, Saya H and Hara E: Mitogenic signalling and the p16INK4a-Rb pathway cooperate to enforce irreversible cellular senescence. Nat Cell Biol 8: 1291-1297, 2006
- Ozawa T, Araki N, Yunoue S, Tokuo H, Feng L, Patrakitkomjorn S, Hara T, Ichikawa Y, Matsumoto K, Fujii K and Saya H: The neurofibromatosis type 1 gene product neurofibromin enhances cell motility by regulating actin filament dynamics via the Rho-ROCK-LIMK2-cofilin pathway. J Biol Chem 280: 39524-39533, 2005
- Nagano O, Murakami D, Hartmann D, De Strooper B, Saftig P, Iwatsubo T, Nakajima M, Shinohara M and Saya H: Cell-matrix interaction via CD44 is independently regulated by different metalloproteinases activated in response to extracellular Ca2+ influx and PKC activation. J Cell Biol 165: 893-902, 2004.
Copyright © Keio University. All rights reserved.
